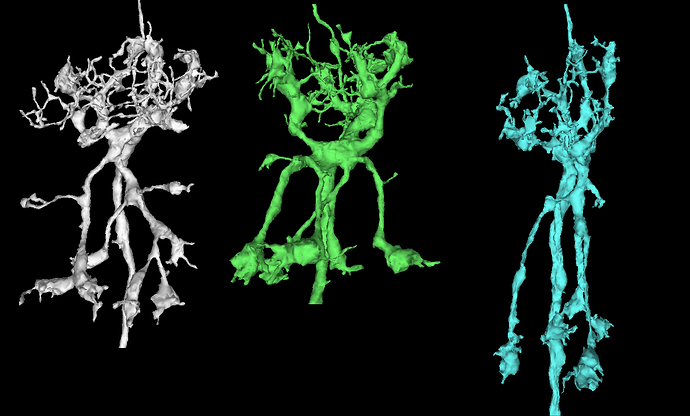I decided to go ahead and create a thread solely devoted to this topic, as it has proven to be a very useful and practical approach to dealing with some of the issues encountered in the EM imaging data. I do realize that most of what is presented below will already be familiar to most of you. But, I felt that it would be worthwhile to document our observations here, in a separate topic, as a reference for ourselves, and as a primer for the benefit of newcomers.
Inferring Microstructure from Macrostructure
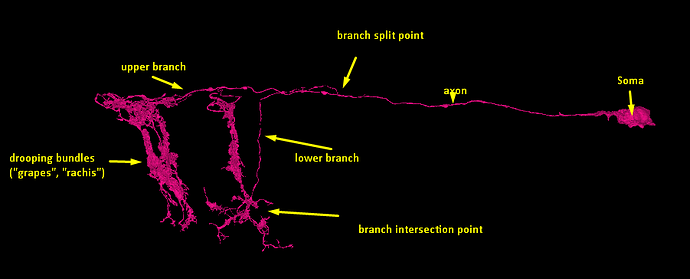
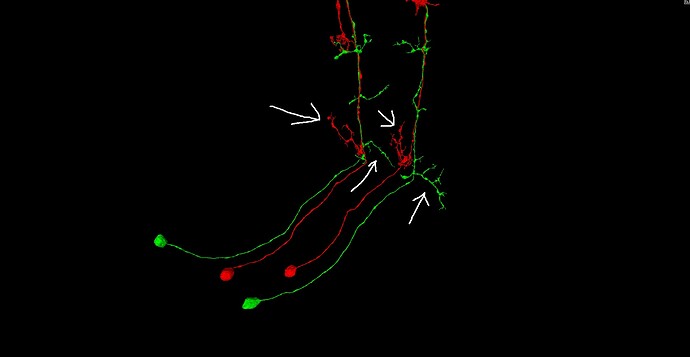
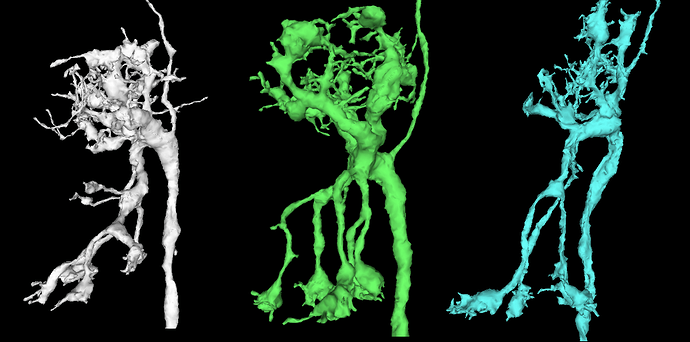
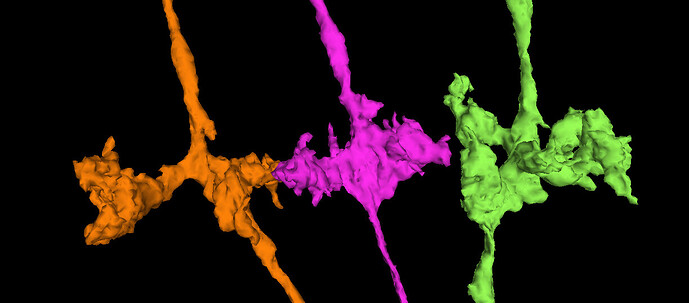
So, what do we mean by “inferring microstructure from macrostructure”? Well, suppose that you own a small orchard, of hypothetical trees that always grow in the same way, and always have three branches. The trees in your orchard are arranged in perfect geometric rows, with each tree being a nearly identical copy of every other tree. Now suppose that you go for a walk in your orchard, one day, and you come across a peculiar tree with only two branches, and find a dead branch laying on the ground beside it. It would be perfectly reasonable to “infer” that the dead branch on the ground is, in fact, the missing third branch of the peculiar tree. This would be an example of “inferring microstructure from macrostructure”.
Columnar Structures and Row Fields
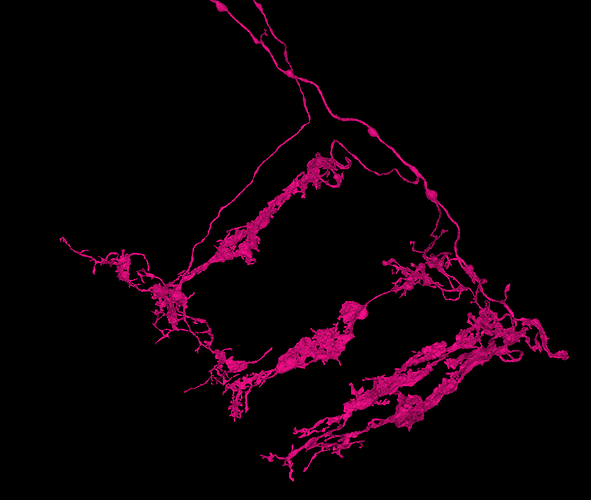
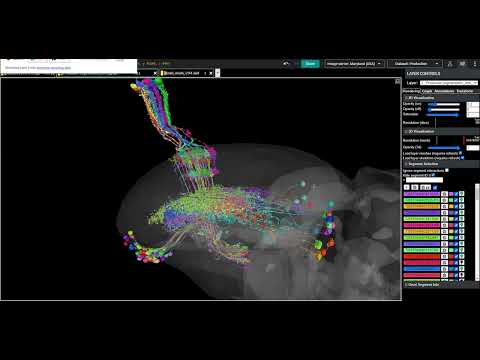
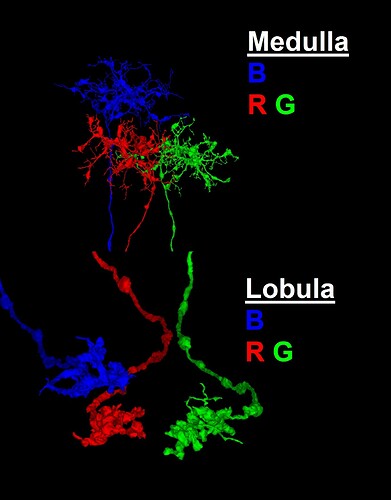
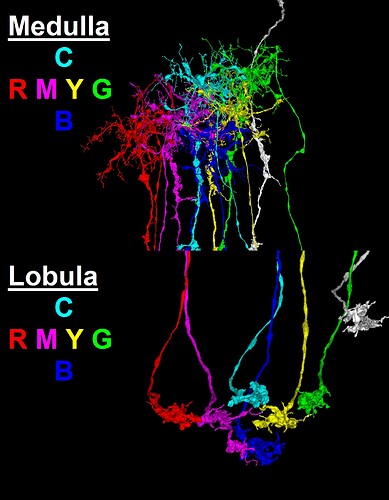
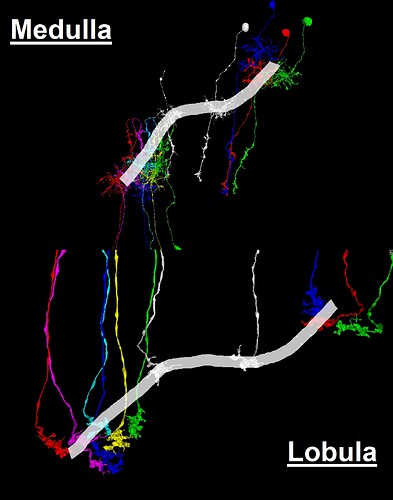
The link pictured below (as a clickable YouTube video thumbnail) contains a collection of neurons selected from the “Zone 2” set (in the Flywire blog), and several additionally selected neurons. This selected group of neurons is presented here as being representative of the general overall structure of the optic lobe. As can be seen, the structure of the optic lobe can be characterized as the combination of an array of regularly spaced and repeated columnar structures, with a series of several successive layers of perpendicular row fields, intersecting the columnar array. A familiar example of the columnar structures of the optic lobe, would be the L cells of the ubiquitous “laminar cartridges” (found repeated throughout the lamina), which represent the “tops” of the columnar structures of the optic lobe. A familiar example of the well defined row fields of the optic lobe, would be the Dm cells (found repeated throughout the medulla), which have been featured in the “Gallery of Amusements” topic, for their aesthetically pleasing appearance, which sometimes resembles a field of flowers.
Applying the above, as an approach…
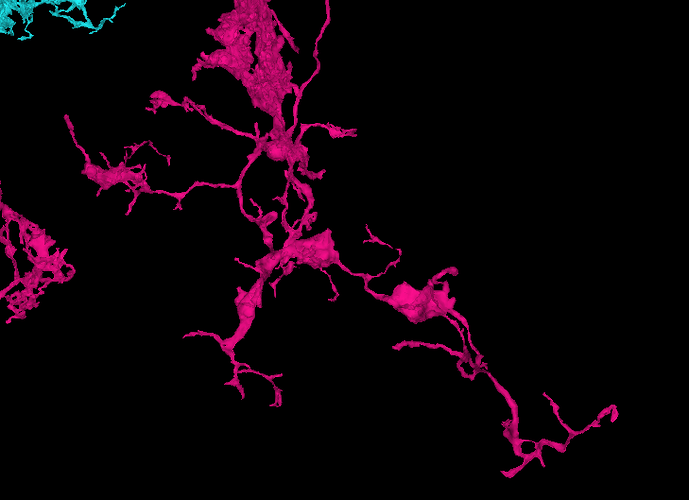
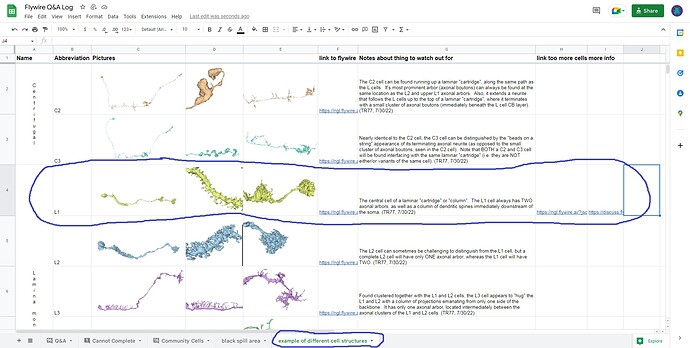
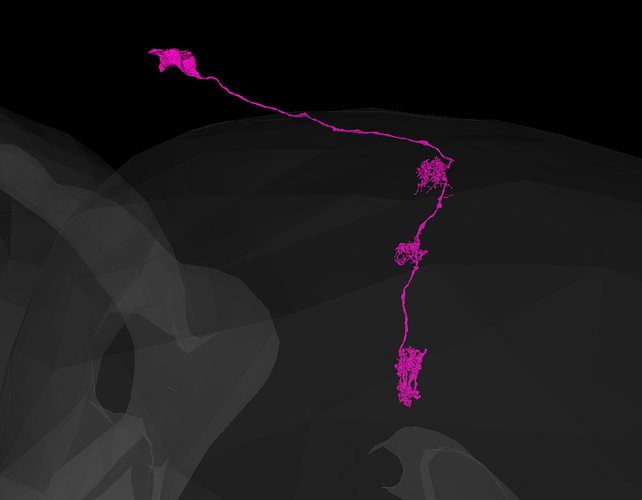
Recognizing the overall general structure of the optic lobe, as well as the component substructures, can be helpful in anticipating where and what to look for, when searching for the extensions of a particular cell, or group of cells. For example, if your attention is called to a particular L cell in the lamina, then without even looking, you can predict it’s soma location, arbor locations, and overall morphology, as well as anticipate that it neighbors and interfaces with several other L cells in a predictable way, along with several R’s, T’s, C’s, etc. This consistency and predictability of structure can be of tremendous value, when problems in the EM imaging data are encountered.
In closing…
Feel free to post here as much as you like, as the search tool can always be used to locate posts relevant to a particular cell type or structure. A “useful structural pattern” could be anything from noticing consistency in the shape of a particular cell type’s arbors, to noticing that a small group of cell types almost always occur together and in the same arrangement, and even noticing large scale structural patterns related to the overall structure of the optic lobe.
Cheers, all.
https://ngl.flywire.ai/?json_url=https://globalv1.flywire-daf.com/nglstate/5451622914195456